library(tximport)
folders <- dir(pattern="SRR21568*")
samples <- sub("_quant", "", folders)
files <- file.path( folders, "abundance.h5" )
names(files) <- samples
txi.kallisto <- tximport(files, type = "kallisto", txOut = TRUE)1 2 3 4 library(tximport)
folders <- dir(pattern="SRR21568*")
samples <- sub("_quant", "", folders)
files <- file.path( folders, "abundance.h5" )
names(files) <- samples
txi.kallisto <- tximport(files, type = "kallisto", txOut = TRUE)1 2 3 4 head(txi.kallisto$counts) SRR2156848 SRR2156849 SRR2156850 SRR2156851
ENST00000539570 0 0 0.00000 0
ENST00000576455 0 0 2.62037 0
ENST00000510508 0 0 0.00000 0
ENST00000474471 1 1 1.00000 0
ENST00000381700 0 0 0.00000 0
ENST00000445946 0 0 0.00000 0colSums(txi.kallisto$counts)SRR2156848 SRR2156849 SRR2156850 SRR2156851
586450 2600800 2372309 2111474 sum(rowSums(txi.kallisto$counts)>0)[1] 94657to.keep <- rowSums(txi.kallisto$counts) > 0
kset.nonzero <- txi.kallisto$counts[to.keep,]keep2 <- apply(kset.nonzero,1,sd)>0
x <- kset.nonzero[keep2,]pca <- prcomp(t(x), scale=TRUE)
summary(pca)Importance of components:
PC1 PC2 PC3 PC4
Standard deviation 201.1698 168.6387 160.4157 0.70709
Proportion of Variance 0.4276 0.3005 0.2719 0.00001
Cumulative Proportion 0.4276 0.7281 1.0000 1.00000plot(pca$x[,1], pca$x[,2],
col=c("blue","blue","red","red"),
xlab="PC1", ylab="PC2", pch=16)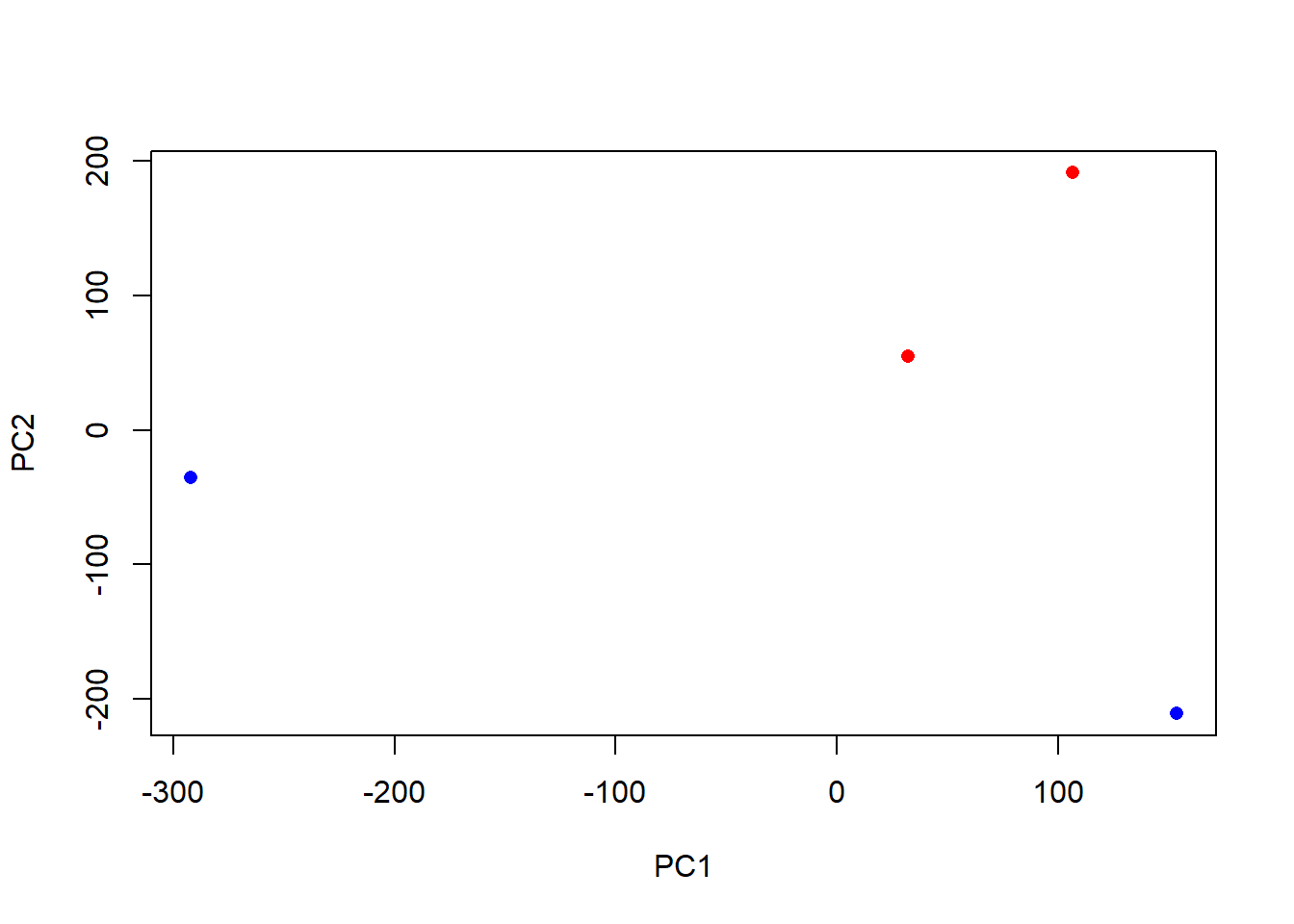
plot(pca$x[,1], pca$x[,3],
col=c("blue","blue","red","red"),
xlab="PC1", ylab="PC3", pch=16)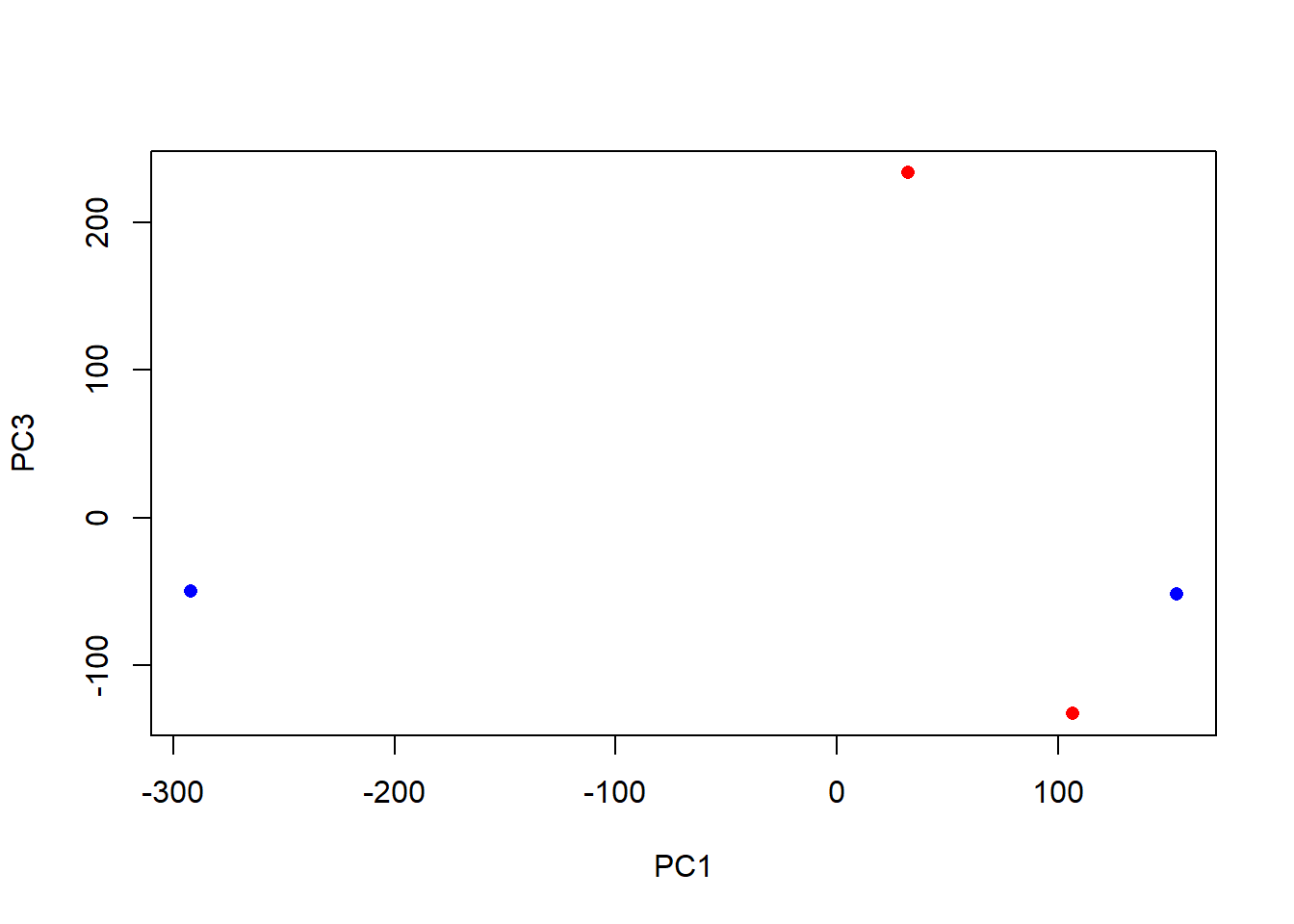
plot(pca$x[,2], pca$x[,3],
col=c("blue","blue","red","red"),
xlab="PC2", ylab="PC3", pch=16)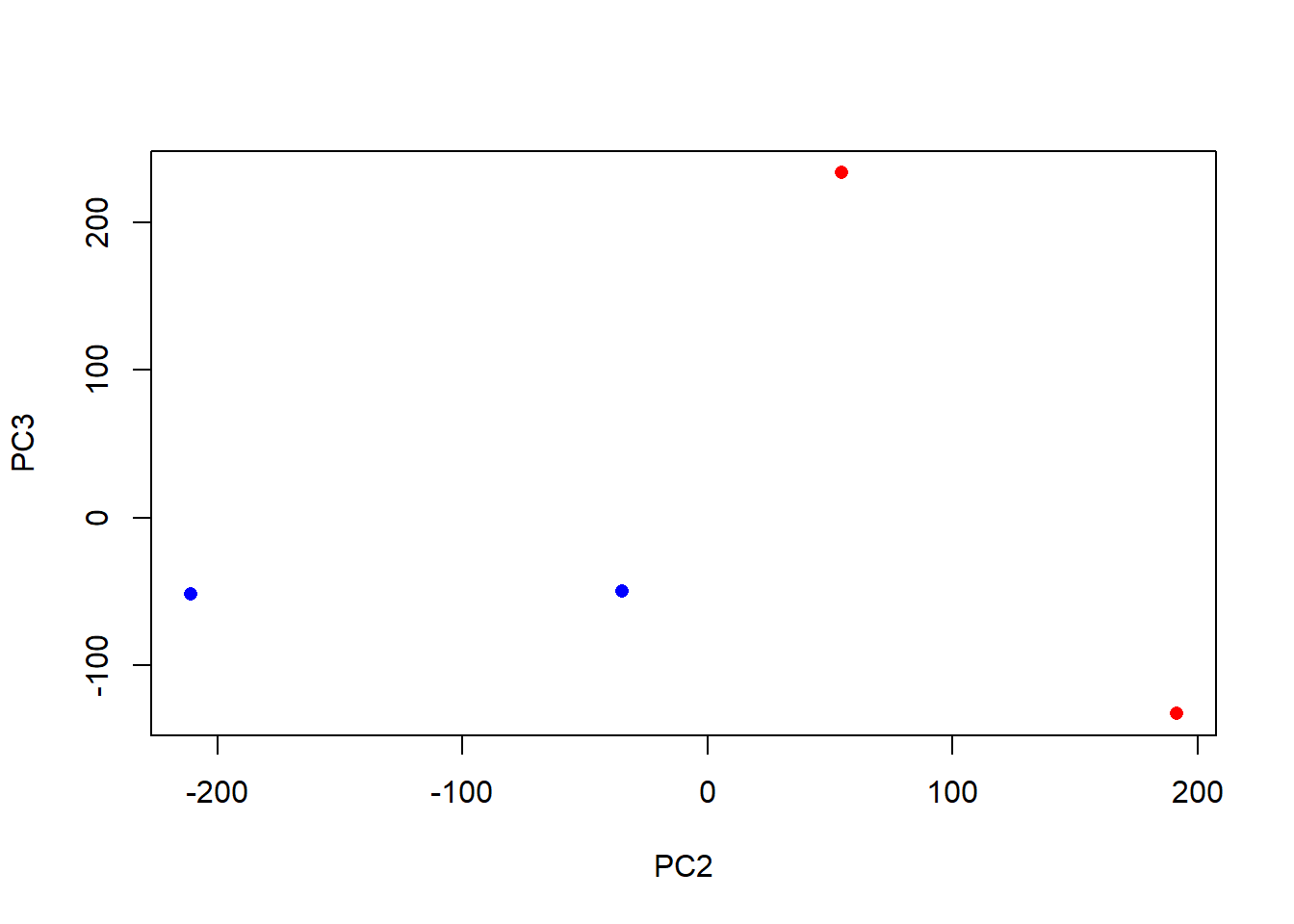
library(ggplot2)
library(ggrepel)
mycols <- c("blue","blue","red","red")
ggplot(pca$x) +
aes(PC1, PC2, label=rownames(pca$x)) +
geom_point( col=mycols ) +
geom_text_repel( col=mycols ) +
theme_bw()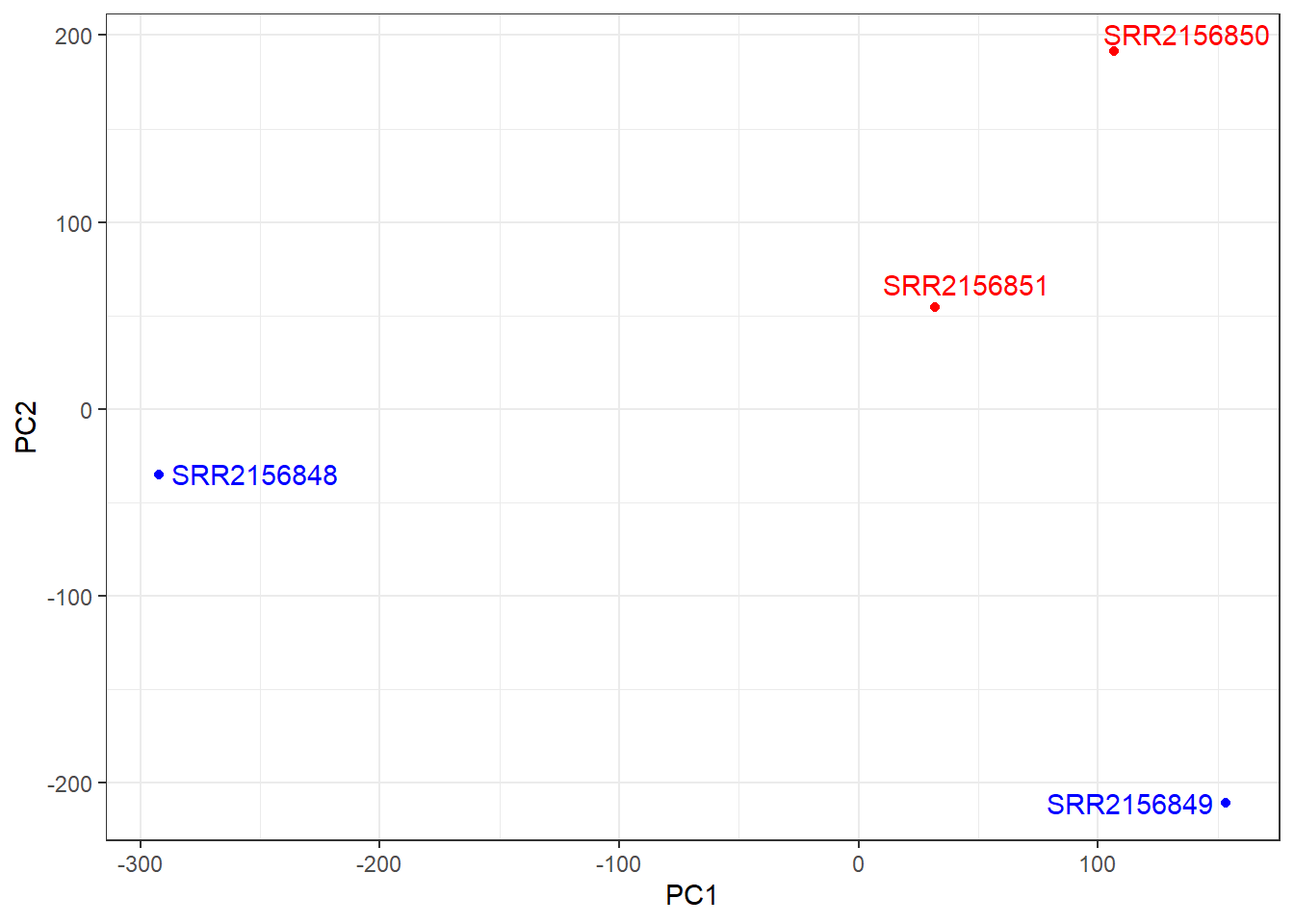
ggplot(pca$x) +
aes(PC1, PC3, label=rownames(pca$x)) +
geom_point(col=mycols) +
geom_text_repel(col=mycols) +
theme_bw()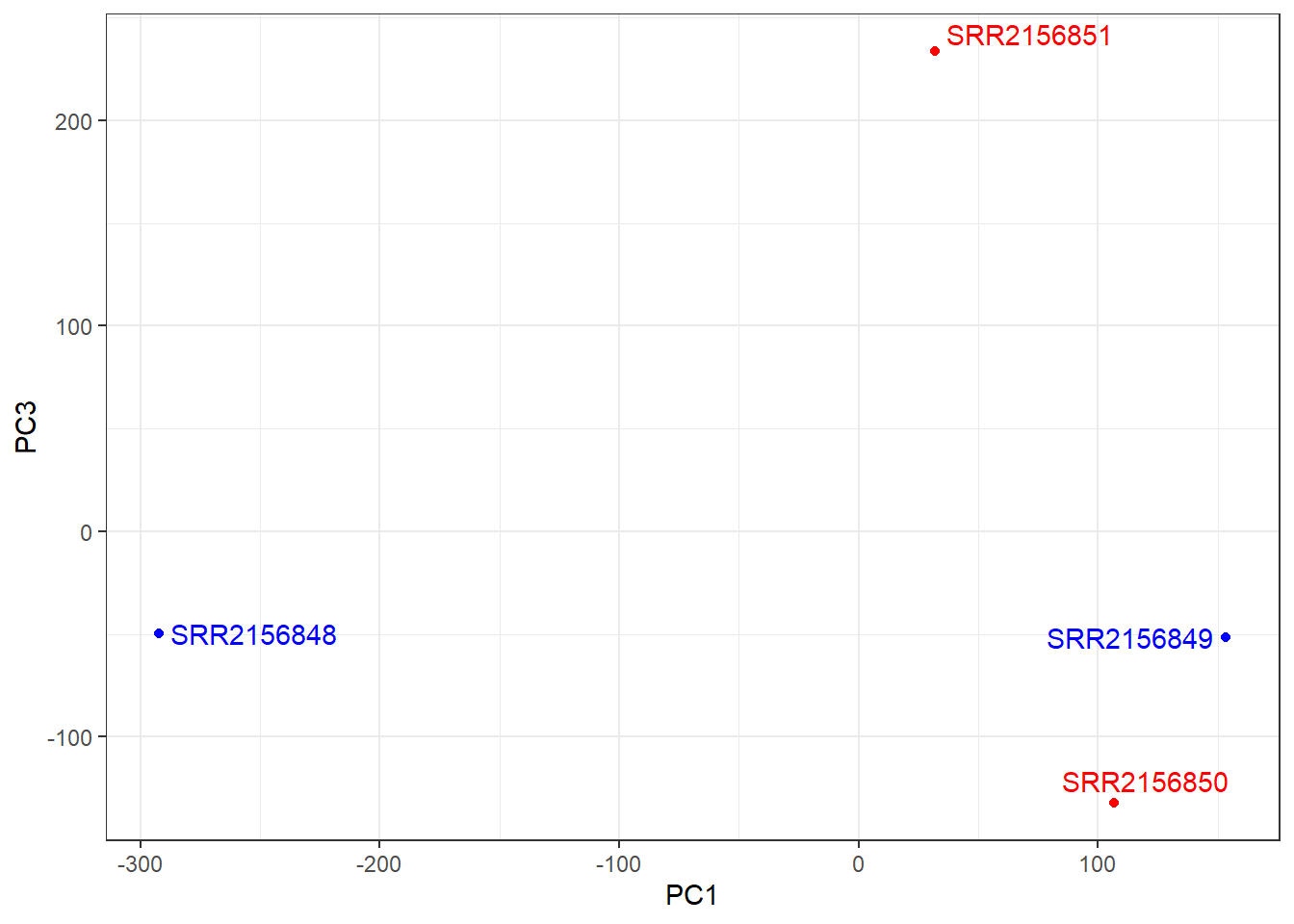
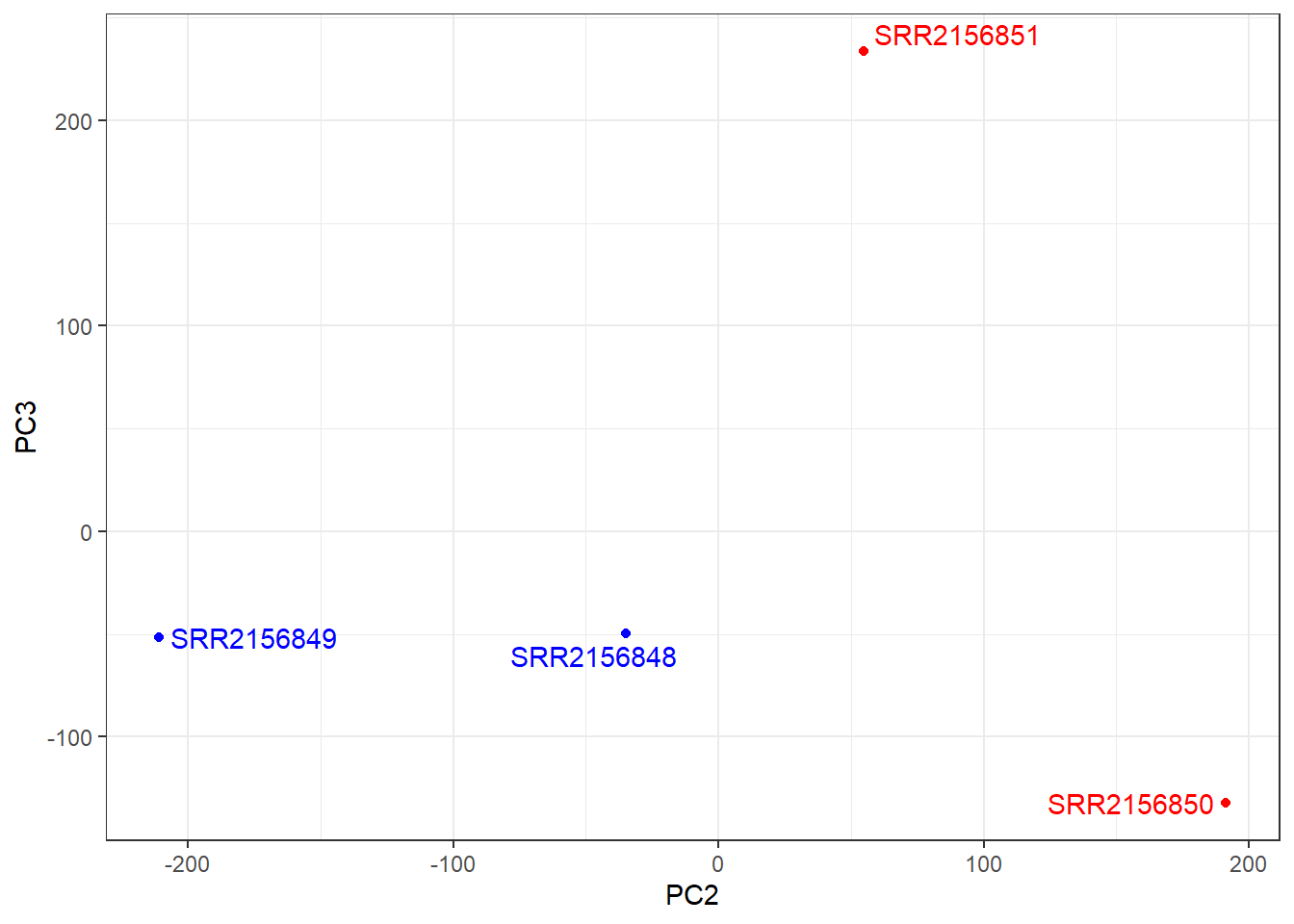
ggplot(pca$x) +
aes(PC2, PC3, label=rownames(pca$x)) +
geom_point(col=mycols) +
geom_text_repel(col=mycols) +
theme_bw()